Dynamics of Soluble HLA-G and the 14-bp Insertion/Deletion Polymorphism as Predictors of Therapeutic Response in Acute Myeloid Leukemia
DOI:
https://doi.org/10.54133/ajms.v10i2.2861Keywords:
Acute myeloid leukemia, Chemotherapy response, Leukemic burden , Soluble HLA-GAbstract
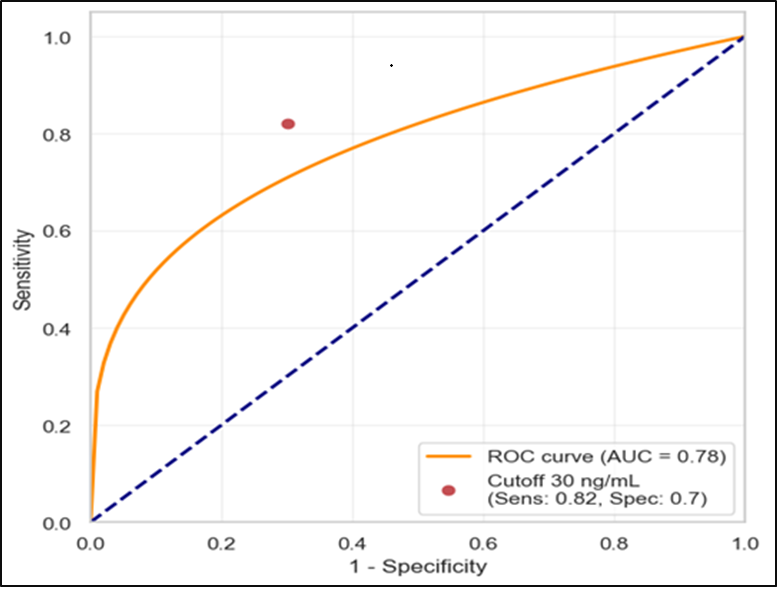
Background: Acute myeloid leukemia (AML) exploits immune escape mechanisms such as human leukocyte antigen-G (HLA-G), a non-classical HLA-I molecule involved in immune checkpoints and associated with the 14-bp insertion/deletion (Ins/Del) polymorphism for cancer prognosis. Prospective data regarding the dynamics of these parameters and their relation to therapy outcomes in AML patients remains limited. Objectives: Determine the relationship between the serum level of soluble human leukocyte antigen-G (sHLA-G) and the 14-bp deletion/insertion (Ins/Del) polymorphism of the sHLA-G gene in AML patients before and after chemotherapy. Methods: A longitudinal study with a prospective follow-up was executed. Forty-six adult patients diagnosed with AML were recruited. ELISA was used to measure serum levels of sHLA-G at diagnosis (T0) and six months after induction chemotherapy (T1). PCR genotyping was done on the HLA-G 14-bp polymorphism. This study focuses on the interaction of sHLA-G dynamics and genotype with the outcomes of clinical response, which could be Complete Remission (CR) or Non-Remission (NR). Results: sHLA-G levels reduced markedly after induction chemotherapy (28.5±15.3ng/mL at T0 vs. 7.6±6.1ng/mL at T1, p<0.001). Baseline sHLA-G values greater than 30ng/mL were significant predictors of non-response to treatment, demonstrating 82% sensitivity and 70% specificity. However, there were no significant variations in the distribution of HLA-G 14-bp genotypes between responders and non-responders (p=0.45). Conclusions: sHLA-G is a sensitive and dynamic biomarker of leukemic load; high baseline levels are significantly associated with non-response to induction therapy. In contrast, the HLA-G 14-bp polymorphism was not identified as an independent predictor of therapeutic response in this cohort.
Downloads
References
Carosella ED, Moreau P, LeMaoult J, Rouas-Freiss N. HLA-G: from biology to clinical benefits. Trends Immunology. 2008;29(3):125-132. doi: 10.1016/j.it.2007.11.005. DOI: https://doi.org/10.1016/j.it.2007.11.005
Hazini A, Fisher K, Seymour L. Deregulation of HLA-I in cancer and its central importance for immunotherapy. J Immunother Cancer. 2021;9(8):e002899. doi: 10.1136/jitc-2021-002899. DOI: https://doi.org/10.1136/jitc-2021-002899
Rizzo R, Audrito V, Vacca P, Rossi D, Brusa D, Stignani M, et al. HLA-G is a component of the chronic lymphocytic leukemia escape repertoire to generate immune suppression: impact of the HLA-G 14 base pair (rs66554220) polymorphism. Haematologica. 2014;99(5):888-896. doi: 10.3324/haematol.2013.095281. DOI: https://doi.org/10.3324/haematol.2013.095281
Loustau M, Anna F, Dréan R, Lecomte M, Langlade-Demoyen P, Caumartin J. HLA-G neo-expression on tumors. Front Immunol. 2020;11:1685. doi: 10.3389/fimmu.2020.01685. DOI: https://doi.org/10.3389/fimmu.2020.01685
Siegel RL, Miller KD, Fuchs HE, Jemal A. Cancer statistics, 2022. CA Cancer J Clin. 2022;72(1):7-33. doi: 10.3322/caac.21708. DOI: https://doi.org/10.3322/caac.21708
Al-Tamimi J, Al Omar SY, Al-Khulaifi F, Alharbi SA, Al-jurayyan A, Mansour L. Evaluation of the relationships between HLA-G 14 bp polymorphism and two acute leukemia in a Saudi population. J King Saud Univ Sci. 2022;34(6):102139. doi: 10.1016/j.jksus.2022.102139. DOI: https://doi.org/10.1016/j.jksus.2022.102139
Arnaiz-Villena A, Juarez I, Suarez-Trujillo F, López-Nares A, Vaquero C, Palacio-Gruber J, et al. HLA-G: Function, polymorphisms and pathology. Int J Immunogenet. 2021;48(2):172-192. doi: 10.1111/iji.12513. DOI: https://doi.org/10.1111/iji.12513
van der Ven K, Pfeiffer K, Skrablin S. HLA-G polymorphisms and molecule function--questions and more questions--a review. Placenta. 2000(Suppl A):S86-92. doi: 10.1053/plac.1999.0515. DOI: https://doi.org/10.1053/plac.1999.0515
Henig I, Zuckerman T. Hematopoietic stem cell transplantation-50 years of evolution and future perspectives. Rambam Maimonides Med J. 2014;5(4):e0028. doi: 10.5041/RMMJ.10162. DOI: https://doi.org/10.5041/RMMJ.10162
Bartolome J, Molto C, Benitez-Fuentes JD, Fernandez-Hinojal G, Manzano A, Perez-Segura P, et al. Prognostic value of human leukocyte antigen G expression in solid tumors: a systematic review and meta-analysis. Front Immunol. 2023;14:1165813. doi: 10.3389/fimmu.2023.1165813. DOI: https://doi.org/10.3389/fimmu.2023.1165813
Celik A. The basis of HLA-G mediated dysregulation of immune effector cells: Dissertation, Hannover, Medizinische Hochschule Hannover, 2018; 2018.
Locafaro G, Amodio G, Tomasoni D, Tresoldi C, Ciceri F, Gregori S. HLA-G expression on blasts and tolerogenic cells in patients affected by acute myeloid leukemia. J Immunol Res. 2014;2014:636292. doi: 10.1155/2014/636292. DOI: https://doi.org/10.1155/2014/636292
Yan WH. HLA-G expression in cancers: potential role in diagnosis, prognosis and therapy. Endocr Metab Immune Disord Drug Targets. 2011;11(1):76-89. doi: 10.2174/187153011794982059. DOI: https://doi.org/10.2174/187153011794982059
Paul P, Cabestre FA, Ibrahim EC, Lefebvre S, Khalil-Daher I, Vazeux G, et al. Identification of HLA-G7 as a new splice variant of the HLA-G mRNA and expression of soluble HLA-G5, -G6, and -G7 transcripts in human transfected cells. Hum Immunol. 2000;61(11):1138-1149. doi: 10.1016/s0198-8859(00)00197-x. DOI: https://doi.org/10.1016/S0198-8859(00)00197-X
Morandi F, Airoldi I. HLA-G and other immune checkpoint molecules as targets for novel combined immunotherapies. Int J Mol Sci. 2022;23(6):2925. doi: 10.3390/ijms23062925. DOI: https://doi.org/10.3390/ijms23062925
Nückel H, Rebmann V, Dürig J, Dührsen U, Grosse-Wilde H. HLA-G expression is associated with an unfavorable outcome and immunodeficiency in chronic lymphocytic leukemia. Blood. 2005;105(4):1694-1698. doi: 10.1182/blood-2004-08-3335. DOI: https://doi.org/10.1182/blood-2004-08-3335
Caocci G, Greco M, Fanni D, Senes G, Littera R, Lai S, et al. HLA-G expression and role in advanced-stage classical Hodgkin lymphoma. Eur J Histochem. 2016;60(2):2606. doi: 10.4081/ejh.2016.2606. DOI: https://doi.org/10.4081/ejh.2016.2606
Tawfeek GA, Alhassanin S. HLA-G gene polymorphism in Egyptian patients with non-Hodgkin lymphoma and its clinical outcome. Immunol Invest. 2018;47(3):315-325. doi: 10.1080/08820139.2018.1430826. DOI: https://doi.org/10.1080/08820139.2018.1430826
Waterhouse M, Duque-Afonso J, Wäsch R, Bertz H, Finke J. Soluble HLA-G molecules and HLA-G 14-base pair polymorphism after allogeneic hematopoietic cell transplantation. Transplant Proc. 2013;45(1):397-401. doi: 10.1016/j.transproceed.2012.05.073. DOI: https://doi.org/10.1016/j.transproceed.2012.05.073
Wang C, Su NW, Hsu K, Kao CW, Chang MC, Chang YF, et al. The implication of serum HLA-G in angiogenesis of multiple myeloma. Mol Med. 2024;30(1):86. doi: 10.1186/s10020-024-00860-5. DOI: https://doi.org/10.1186/s10020-024-00860-5
Moreau P, Adrian-Cabestre F, Menier C, Guiard V, Gourand L, Dausset J, et al. IL-10 selectively induces HLA-G expression in human trophoblasts and monocytes. Int Immunol. 1999;11(5):803-811. doi: 10.1093/intimm/11.5.803. DOI: https://doi.org/10.1093/intimm/11.5.803
Downloads
Published
How to Cite
Issue
Section
License
Copyright (c) 2026 Al-Rafidain Journal of Medical Sciences ( ISSN 2789-3219 )

This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License.
Published by Al-Rafidain University College. This is an open access journal issued under the CC BY-NC-SA 4.0 license (https://creativecommons.org/licenses/by-nc-sa/4.0/).











